Ancient DNA Clovis Anzick and HAM DNA Group #1
Because of the destroyed or missing records in Virginia, I had been working on the autosomal Half Life equation in order to tie our group to Somerset by use of autosomal DNA. Previously, we had seen that two kits from Somerset, England are a match to HAM DNA Group #1 (I1-M253).
I has recently noticed that the Half Life equation was throwing off errors, or variation from what one would expect to see from the Half Life equation. This was particularly troublesome when the SNP density ratio was between 1.0 and 1.5 where RATIO = SNPs/(100*cMs).
So, when I ran across an article on Ancient DNA by Roberta Estes, I became curious as to what the half Life equation might look like when used on Ancient DNA.
"Analyzing the Native American Clovis Anzick Ancient Results" DNAeXplained – Genetic Genealogy
Roberta had been talking to Felix Chandrakamur about the Clovis results that had been uploaded to GEDMatch, and Roberta had noticed that the Clovis upload was matching living people. She had found 1466 matches to Clovis at GEDMatch above the 7 cM level.
This is a stunning result. For a little background on the Half Life equation, you can find it's limit by plugging "1 cM," which shows that the equation is built to display a result of about 8th cousin level (or 9 generations) for a shared segment of 1 cM in size.
Half Life = - LN(1/281.5)/.693147
Half Life = 8.1
For the 11th cousin level, it needs a segment size of 0.1 cMs.
Clearly, if the Clovis sample is 12,500 years old and is matching living people at 7 cMs and above, then the Half Life equation is useless in it's current form.
Now, the usual argument might be that these Clovis samples are Identical by State (IBS), and not Identical by Descent (IBD). For example, the current ISOGG statistics show that the smallest shared segment equivalent to 5th cousins is 3.32 cMs.
See:
"Autosomal DNA statistics" at ISOGG
Or see also:
"Cousin statistics" at ISOGG
See the scientific paper for the expected (i.e., theoretical) number of cMs at the 5th cousin level:
"Cryptic Distant Relatives Are Common in Both Isolated and Cosmopolitan Genetic Samples"
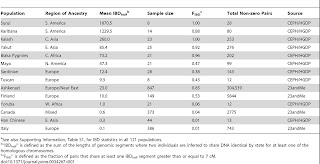 | |
| Table 1. Expected extent of IBD and number of cousins for 1st–10th degrees of cousinship. https://doi.org/10.1371/journal.pone.0034267.t002 |
Also, it is instructive to note that a good quality 10 cM segment was extracted from the Altai Neanderthal who lived 50,000 years ago in Siberia.
- see "The complete genome sequence of a Neanderthal from the Altai Mountains"
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4031459/
"To estimate the extent of their relatedness, we scanned the genome for 1Mb regions where most non-overlapping 50-kb-windows were devoid of heterozygous sites and merged adjacent regions (SI 10). The Neandertal genome has 20 such regions longer than 10cM whereas the Denisovan genome has one."
Finally, Roberta Estes wrote an article regarding possible sampling errors, due to the nature of conversion for upload to GEDMatch. The basic concern appears to be "No-Call" rates:
"Ancient DNA Matching – A Cautionary Tale" DNAeXplained – Genetic Genealogy, by Roberta Estes
Roberta also explains that subsequent comparisons do not match previous comparisons. I have seen that with GEDMatch data, particularly when he changes versions of the 'One to One' Utility (which is now at version 2.1.1(c). I have had to re-work the data for Group #1 several times over the years, just to keep the data consistent with the current version of the GEDMatch 'One to One' Utility.
Also note that these Clovis segments for kit F999919 match to 12 cMs on living Native Americans.
With that, the first thing I wanted to know was to take a look at the SNP density RATIO for the Clovis comparisons. How do ancient Clovis segments compare with SNP density RATIO, and do these ancient segments also throw off variations from the Half Life equations?
RATIO = SNPs/(100*cMs)
I took a look at hr matching segments for HAM DNA Group #1, plus our 'control group' kit, Arnold (22 kits). I found that out of the seven largest segments matching Clovis, all but five had a RATIO of less than 2.0, or about 3.2%.
To put that into perspective, 7 x 22 = 154 segments
5 segments/154 segments = 3.2%
That means, for our sample (the Ancient Clovis DNA matching shared segments), 97% have an SNP density RATIO of less than 2.0
Among the Clovis shared segments for the group, there are about 8% that are in 'Excess IBD' regions.
Below is a summary table of the results.
 |
| Clovis Largest Shared Segment and HAM DNA Group01 |
The kit in HAM DNA Group #1 with the largest matching segment to Clovis:
A404xxx at 5.9 cMs
- Kits with matching Clovis segments with the largest matching starting and/or ending locations:
Clovis Kit Chr Start Location End Location cMs SNPs
F99919 and T074xxx 1 1,751,874 3,003,550 4.2 261
F99919 and T133xxx 1 1,751,874 3,003,550 4.2 258
F99919 and T630xxx 9 38,523,004 70,819,104 4.0 286
F99919 and T682xxx 9 38,694,680 70,536,108 3.4 148
F99919 and A561xxx 12 11,840,131 12,861,007 3.8 347
F99919 and A832xxx 12 11,840,131 12,861,007 3.8 350
F99919 and A438xxx 17 13,813,353 14,461,941 3.6 251
F99919 and T368xxx 17 13,785,798 14,618,990 4.7 354
F99919 and A984xxx 19 8,282,431 9,460,034 4.2 292
F99919 and T611xxx 19 8,214,446 9,934,324 5.5 401
I would think that these matching Clovis segments would imply a Native American ancestor for the above kits (Amelia County, Ashe County, and the Arkansas lines).
This also implies that if kit A171xxx (of Somerset) does not have a Native American ancestor, then the timeline to connection could be considerably further back in time and may have an impact upon how the Half Life equation should function.
For example, a person in Somerset with a documented line may have a very small chance of having a Native American ancestor. A Clovis match would push the age back at least 12,500 years, much more than the 9 generations from the current 1 cM limit of the Half Life equation.
Sample Location GEDMatch Sex Y-DNA Mt-DNA Approx. Age by authors Felix Chandrakumar Analysis or Comments
Clovis-Anzick-1 Montana, North America F999919 M Q-Z780 D4h3a 12,500 years Matches Living people.
http://www.y-str.org/2014/09/clovis-anzick-dna.html
References:
"Analyzing the Native American Clovis Anzick Ancient Results" DNAeXplained – Genetic Genealogy, Roberta Estes
https://dna-explained.com/2014/09/23/analyzing-the-native-american-clovis-anzick-ancient-results/
"Ancient DNA Matching – A Cautionary Tale" DNAeXplained – Genetic Genealogy, by Roberta Estes
https://dna-explained.com/2014/09/30/ancient-dna-matching-a-cautionary-tale/
"Matching DNA of Living Native Descendants to DNA of Native Ancestors"
https://nativeheritageproject.com/2014/09/25/matching-dna-of-living-native-descendants-to-dna-of-native-ancestors/
"Autosomal DNA statistics"
https://isogg.org/wiki/Autosomal_DNA_statistics
Or see also:
"Cousin statistics"
https://isogg.org/wiki/Cousin_statistics
The scientific paper for the expected (i.e., theoretical) number of cMs at the 5th cousin level:
"Cryptic Distant Relatives Are Common in Both Isolated and Cosmopolitan Genetic Samples"
https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0034267
GEDMatch John Olson depends upon financial contributions.
Clovis-Anzick-1 Montana, North America F999919 M Q-Z780 D4h3a 12,500 years Matches Living people.
http://www.y-str.org/2014/09/clovis-anzick-dna.html
Autosomal DNA Half Life Equation
http://hamcountry-blog.blogspot.com/2018/02/autosomal-dna-half-life-equation.html